Description
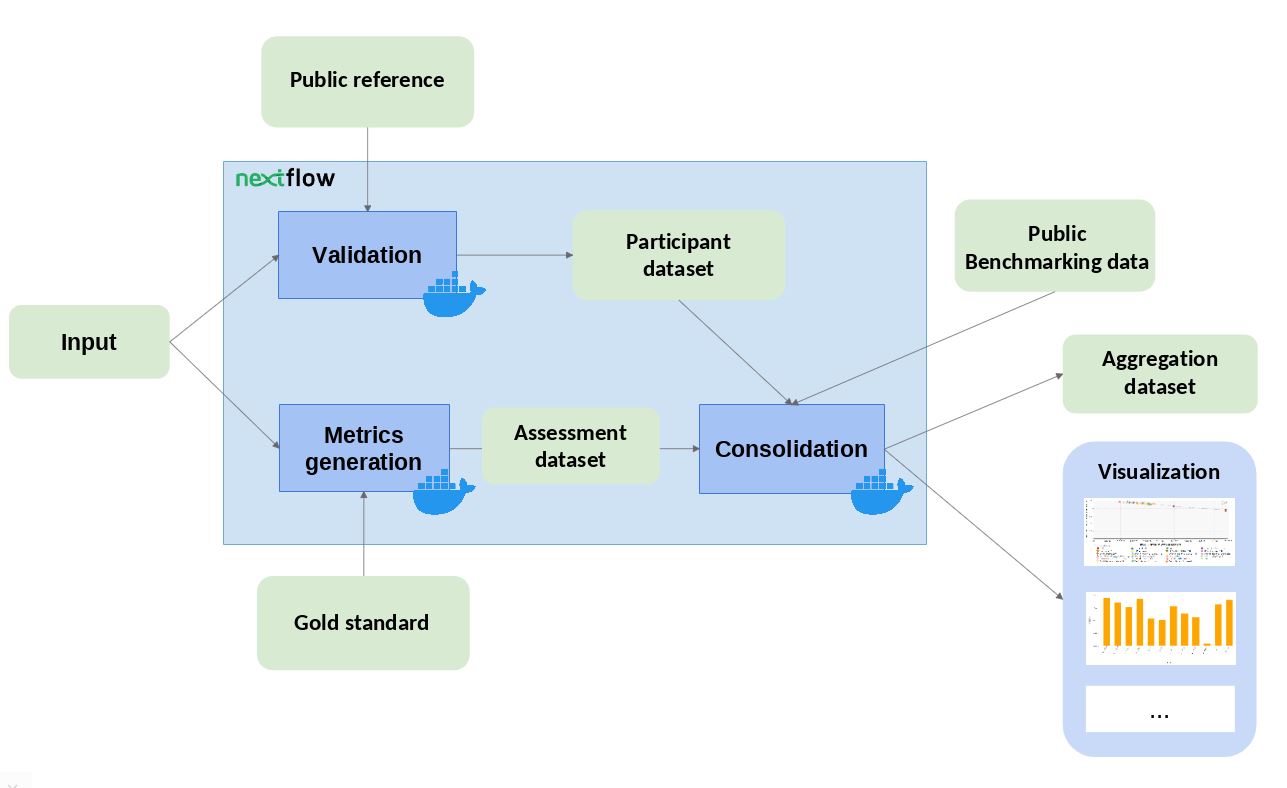
The workflow takes an input file with Cancer Driver Genes predictions (i.e. the results provided by a participant), computes a set of metrics, and compares them against the data currently stored in OpenEBench within the TCGA community. Two assessment metrics are provided for that predictions. Also, some plots (which are optional) that allow to visualize the performance of the tool are generated. The workflow consists in three standard steps, defined by OpenEBench. The tools needed to run these steps are containerised in three Docker images, whose recipes are available in the TCGA_benchmarking_dockers repository and the images are stored in the INB GitLab container registry . Separated instances are spawned from these images for each step:
- Validation: the input file format is checked and, if required, the content of the file is validated (e.g check whether the submitted gene IDs exist)
- Metrics Generation: the predictions are compared with the 'Gold Standards' provided by the community, which results in two performance metrics - precision (Positive Predictive Value) and recall(True Positive Rate).
- Consolidation: the benchmark itself is performed by merging the tool metrics with the rest of TCGA data. The results are provided in JSON format and SVG format (scatter plot).
Data
- TCGA_sample_data folder contains all the reference data required by the steps. It is derived from the manuscript:
Comprehensive Characterization of Cancer Driver Genes and Mutations, Bailey et al, 2018, Cell
- TCGA_sample_out folder contains an example output for a worklow run, with two cancer types / challenges selected (ACC, BRCA). Results obtained from the default execution should be similar to those ones available in this directory. Results found in TCGA_sample_out/results can be visualized in the browser using
benchmarking_workflows_results_visualizerjavascript library.
Requirements
This workflow depends on three tools that have to be installed before you can run it:
- Git: Used to download the workflow from GitHub.
- Docker: The Docker Engine is used under the hood to execute the containerised steps of the benchmarking workflow.
- Nextflow: Is the technology used to write and execute the benchmarking workflow. Note that it depends on Bash (>=3.2) and Java (>=8 , <=17). We provide the script run_local_nextflow.bash that automates their installation for local testing.
Check that these tools are available in your environment:
# Git
> which git
/usr/bin/git
> git --version
git version 2.26.2
# Docker
> which docker
/usr/bin/docker
> docker --version
Docker version 20.10.9-ce, build 79ea9d308018
# Nextflow
> which nextflow
/home/myuser/bin/nextflow
> nextflow -version
N E X T F L O W
version 21.04.1 build 5556
created 14-05-2021 15:20 UTC (17:20 CEST)
cite doi:10.1038/nbt.3820
http://nextflow.io
In the case of docker, apart from being installed the daemon has to be running. On Linux distributions that use Systemd for service management, which includes the most popular ones as of 2021 (Ubuntu, Debian, CentOs, Red Hat, OpenSuse), the systemctl command can be used to check its status and manage it:
# Check status of docker daemon
> sudo systemctl status docker
● docker.service - Docker Application Container Engine
Loaded: loaded (/usr/lib/systemd/system/docker.service; disabled; vendor preset: disabled)
Active: inactive (dead)
Docs: http://docs.docker.com
# Start docker daemon
> sudo systemctl start docker
Download workflow
Simply clone the repository and check out the latest tag (currently 1.0.8):
# Clone repository
> git clone https://github.com/inab/TCGA_benchmarking_dockers.git
# Move to new directory
cd TCGA_benchmarking_workflow/
# Checkout version 1.0.8
> git checkout 1.0.8 -b 1.0.8
Usage
The workflow can be run workflow in two different ways:
- Standard:
nextflow run main.nf -profile docker - Using the bash script that installs Java and Nextflow:
./run_local_nextflow.bash run main.nf -profile docker.
Arguments specifications:
Usage:
Run the pipeline with default parameters:
nextflow run main.nf -profile docker
Run with user parameters:
nextflow run main.nf -profile docker --predictionsFile {driver.genes.file} --public_ref_dir {validation.reference.file} --participant_name {tool.name} --metrics_ref_dir {gold.standards.dir} --cancer_types {analyzed.cancer.types} --assess_dir {benchmark.data.dir} --results_dir {output.dir}
Mandatory arguments:
--input List of cancer genes prediction
--community_id Name or OEB permanent ID for the benchmarking community
--public_ref_dir Directory with list of cancer genes used to validate the predictions
--participant_id Name of the tool used for prediction
--goldstandard_dir Dir that contains metrics reference datasets for all cancer types
--challenges_ids List of types of cancer selected by the user, separated by spaces
--assess_dir Dir where the data for the benchmark are stored
Other options:
--validation_result The output directory where the results from validation step will be saved
--augmented_assess_dir Dir where the augmented data for the benchmark are stored
--assessment_results The output directory where the results from the computed metrics step will be saved
--outdir The output directory where the consolidation of the benchmark will be saved
--statsdir The output directory with nextflow statistics
--data_model_export_dir The output dir where json file with benchmarking data model contents will be saved
--otherdir The output directory where custom results will be saved (no directory inside)
Flags:
--help Display this message
Default input parameters and Docker images to use for each step can be specified in the config file.
NOTE: In order to make your workflow compatible with the OpenEBench VRE Nextflow Executor, please make sure to use the same parameter names in your workflow.
Version History
Version 4 (latest) Created 29th Nov 2021 at 15:21 by Laura Rodriguez-Navas
updated repository tag from 1.0.7 to 1.0.8
Open
 master
masterf6d3a16
Version 3 Created 25th Nov 2021 at 13:31 by Laura Rodriguez-Navas
updated repository tag from 1.0.6 to 1.0.7
Frozen
 master
mastere17c7bd
Version 2 Created 24th Nov 2021 at 14:03 by Laura Rodriguez-Navas
updated repository tag from 1.0.4 to 1.0.6
Frozen
 master
master0bbd457
Version 1 (earliest) Created 23rd Nov 2021 at 16:55 by Laura Rodriguez-Navas
Added/updated 1 files
Frozen
 master
master3a13322
 Creators and Submitter
Creators and SubmitterCreators
Additional credit
Javier Garrayo-Ventas
Submitter
Views: 8198 Downloads: 2792
Created: 23rd Nov 2021 at 16:55
Last updated: 29th Nov 2021 at 15:41
 Tags
Tags Attributions
AttributionsNone
 View on GitHub
View on GitHub
 https://orcid.org/0000-0002-4806-5140
https://orcid.org/0000-0002-4806-5140