Workflow Type: Galaxy
Open
Frozen
Stable
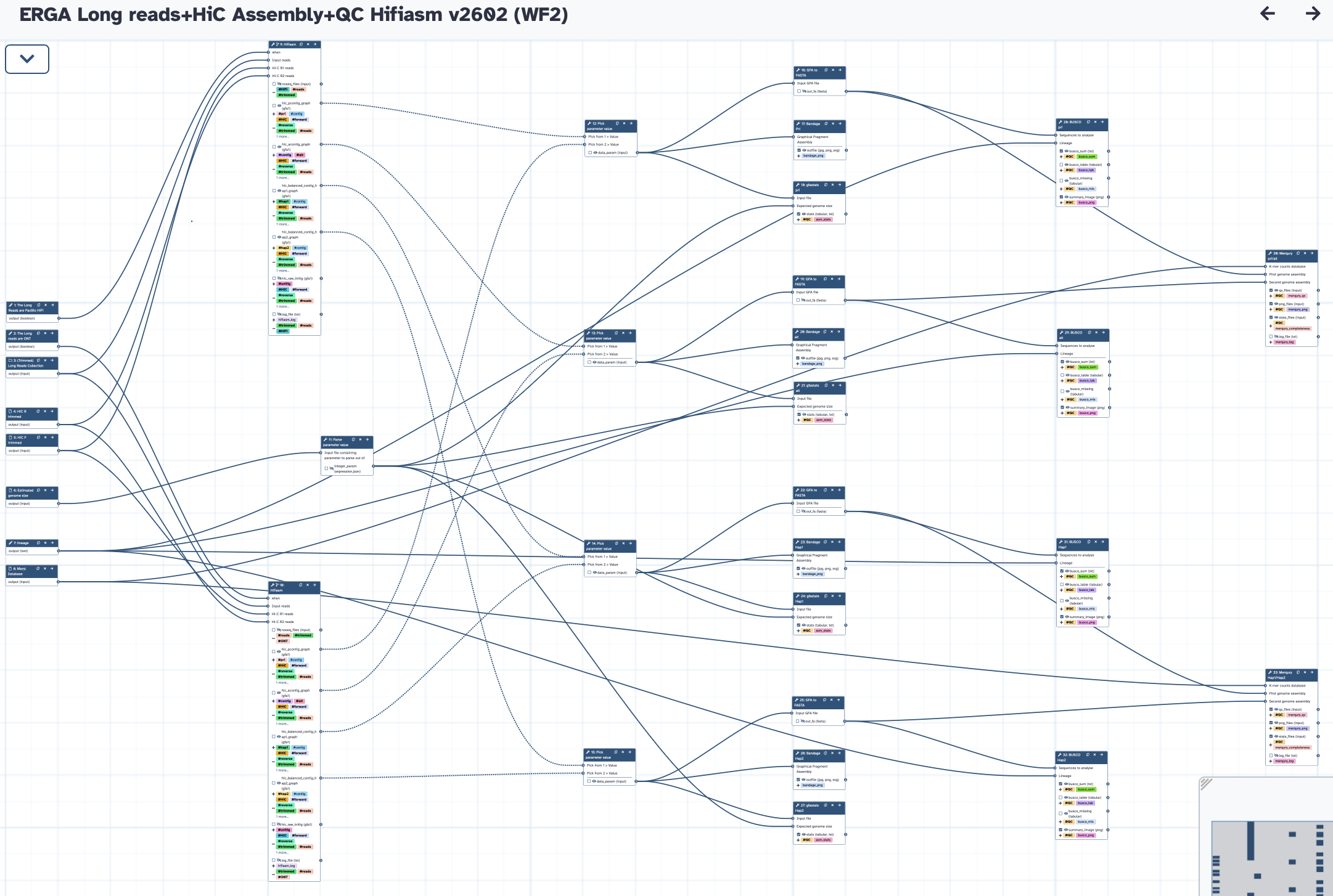
The workflow takes a trimmed long reads collection, and Forward/Reverse HiC reads to run Hifiasm in HiC phasing mode. It produces both Pri/Alt and Hap1/Hap2 assemblies, and runs all the QC analysis (gfastats, BUSCO, and Merqury). The default Hifiasm purge level is aggressive (l3).
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| (Trimmed) Long Reads Collection | (Trimmed) Long Reads Collection | Collection of Long reads in fastq format |
|
| Estimated genome size | Estimated genome size | Select the est_genome_size result obtained during profiling |
|
| HiC F trimmed | HiC F trimmed | n/a |
|
| HiC R trimmed | HiC R trimmed | n/a |
|
| Meryl Database | Meryl Database | Select the meryl_db result obtained during profiling |
|
| The Long Reads are PacBio HiFi | The Long Reads are PacBio HiFi | If your long reads are HiFi, leave it 'yes' |
|
| The Long reads are ONT | The Long reads are ONT | If your long reads are ONT, switch to 'yes'. IMPORTANT: Remember to also switch HiFi to 'no' |
|
| lineage | lineage | lineage for BUSCO, e.g.: arthropoda_odb10, vertebrata_odb10, mammalia_odb10, aves_odb10, tetrapoda_odb10 ... |
|
Steps
| ID | Name | Description |
|---|---|---|
| 8 | Hifiasm | toolshed.g2.bx.psu.edu/repos/bgruening/hifiasm/hifiasm/0.25.0+galaxy1 |
| 9 | Hifiasm | toolshed.g2.bx.psu.edu/repos/bgruening/hifiasm/hifiasm/0.25.0+galaxy1 |
| 10 | Parse parameter value | param_value_from_file |
| 11 | Pick parameter value | toolshed.g2.bx.psu.edu/repos/iuc/pick_value/pick_value/0.2.0 |
| 12 | Pick parameter value | toolshed.g2.bx.psu.edu/repos/iuc/pick_value/pick_value/0.2.0 |
| 13 | Pick parameter value | toolshed.g2.bx.psu.edu/repos/iuc/pick_value/pick_value/0.2.0 |
| 14 | Pick parameter value | toolshed.g2.bx.psu.edu/repos/iuc/pick_value/pick_value/0.2.0 |
| 15 | GFA to FASTA | toolshed.g2.bx.psu.edu/repos/iuc/gfa_to_fa/gfa_to_fa/0.1.2 |
| 16 | Bandage Pri | toolshed.g2.bx.psu.edu/repos/iuc/bandage/bandage_image/2022.09+galaxy4 |
| 17 | gfastats pri | toolshed.g2.bx.psu.edu/repos/bgruening/gfastats/gfastats/1.3.11+galaxy0 |
| 18 | GFA to FASTA | toolshed.g2.bx.psu.edu/repos/iuc/gfa_to_fa/gfa_to_fa/0.1.2 |
| 19 | Bandage alt | toolshed.g2.bx.psu.edu/repos/iuc/bandage/bandage_image/2022.09+galaxy4 |
| 20 | gfastats alt | toolshed.g2.bx.psu.edu/repos/bgruening/gfastats/gfastats/1.3.11+galaxy0 |
| 21 | GFA to FASTA | toolshed.g2.bx.psu.edu/repos/iuc/gfa_to_fa/gfa_to_fa/0.1.2 |
| 22 | Bandage Hap1 | toolshed.g2.bx.psu.edu/repos/iuc/bandage/bandage_image/2022.09+galaxy4 |
| 23 | gfastats Hap1 | toolshed.g2.bx.psu.edu/repos/bgruening/gfastats/gfastats/1.3.11+galaxy0 |
| 24 | GFA to FASTA | toolshed.g2.bx.psu.edu/repos/iuc/gfa_to_fa/gfa_to_fa/0.1.2 |
| 25 | Bandage Hap2 | toolshed.g2.bx.psu.edu/repos/iuc/bandage/bandage_image/2022.09+galaxy4 |
| 26 | gfastats Hap2 | toolshed.g2.bx.psu.edu/repos/bgruening/gfastats/gfastats/1.3.11+galaxy0 |
| 27 | BUSCO pri | toolshed.g2.bx.psu.edu/repos/iuc/busco/busco/5.8.0+galaxy2 |
| 28 | BUSCO alt | toolshed.g2.bx.psu.edu/repos/iuc/busco/busco/5.8.0+galaxy2 |
| 29 | Merqury pri/alt | toolshed.g2.bx.psu.edu/repos/iuc/merqury/merqury/1.3+galaxy4 |
| 30 | BUSCO Hap1 | toolshed.g2.bx.psu.edu/repos/iuc/busco/busco/5.8.0+galaxy2 |
| 31 | BUSCO Hap2 | toolshed.g2.bx.psu.edu/repos/iuc/busco/busco/5.8.0+galaxy2 |
| 32 | Merqury Hap1/Hap2 | toolshed.g2.bx.psu.edu/repos/iuc/merqury/merqury/1.3+galaxy4 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| _anonymous_output_1 | _anonymous_output_1 | n/a |
|
| _anonymous_output_2 | _anonymous_output_2 | n/a |
|
| _anonymous_output_3 | _anonymous_output_3 | n/a |
|
| _anonymous_output_4 | _anonymous_output_4 | n/a |
|
| _anonymous_output_5 | _anonymous_output_5 | n/a |
|
| _anonymous_output_6 | _anonymous_output_6 | n/a |
|
| _anonymous_output_7 | _anonymous_output_7 | n/a |
|
| _anonymous_output_8 | _anonymous_output_8 | n/a |
|
| _anonymous_output_9 | _anonymous_output_9 | n/a |
|
| _anonymous_output_10 | _anonymous_output_10 | n/a |
|
| _anonymous_output_11 | _anonymous_output_11 | n/a |
|
| _anonymous_output_12 | _anonymous_output_12 | n/a |
|
| _anonymous_output_13 | _anonymous_output_13 | n/a |
|
| _anonymous_output_14 | _anonymous_output_14 | n/a |
|
| _anonymous_output_15 | _anonymous_output_15 | n/a |
|
| _anonymous_output_16 | _anonymous_output_16 | n/a |
|
| _anonymous_output_17 | _anonymous_output_17 | n/a |
|
| _anonymous_output_18 | _anonymous_output_18 | n/a |
|
| _anonymous_output_19 | _anonymous_output_19 | n/a |
|
| _anonymous_output_20 | _anonymous_output_20 | n/a |
|
| _anonymous_output_21 | _anonymous_output_21 | n/a |
|
| _anonymous_output_22 | _anonymous_output_22 | n/a |
|
Version History
Version 2 (latest) Created 11th Feb 2026 at 20:03 by Diego De Panis
No revision comments
Open
 master
master63c4f4b
Version 1 (earliest) Created 9th Oct 2023 at 13:47 by Diego De Panis
Initial commit
Frozen
 Version-1
Version-1958e932
 Creators and Submitter
Creators and SubmitterCreator
Additional credit
ERGA
Submitter
Discussion Channel
License
Activity
Views: 8337 Downloads: 749 Runs: 13
Created: 9th Oct 2023 at 13:47
Last updated: 11th Feb 2026 at 20:05
Annotated Properties
 Tags
Tags Attributions
AttributionsNone
 Collections
Collections View on GitHub
View on GitHub Run on Galaxy
Run on Galaxy https://orcid.org/0000-0002-3679-9585
https://orcid.org/0000-0002-3679-9585