Workflow Type: Galaxy
Frozen
Work-in-progress
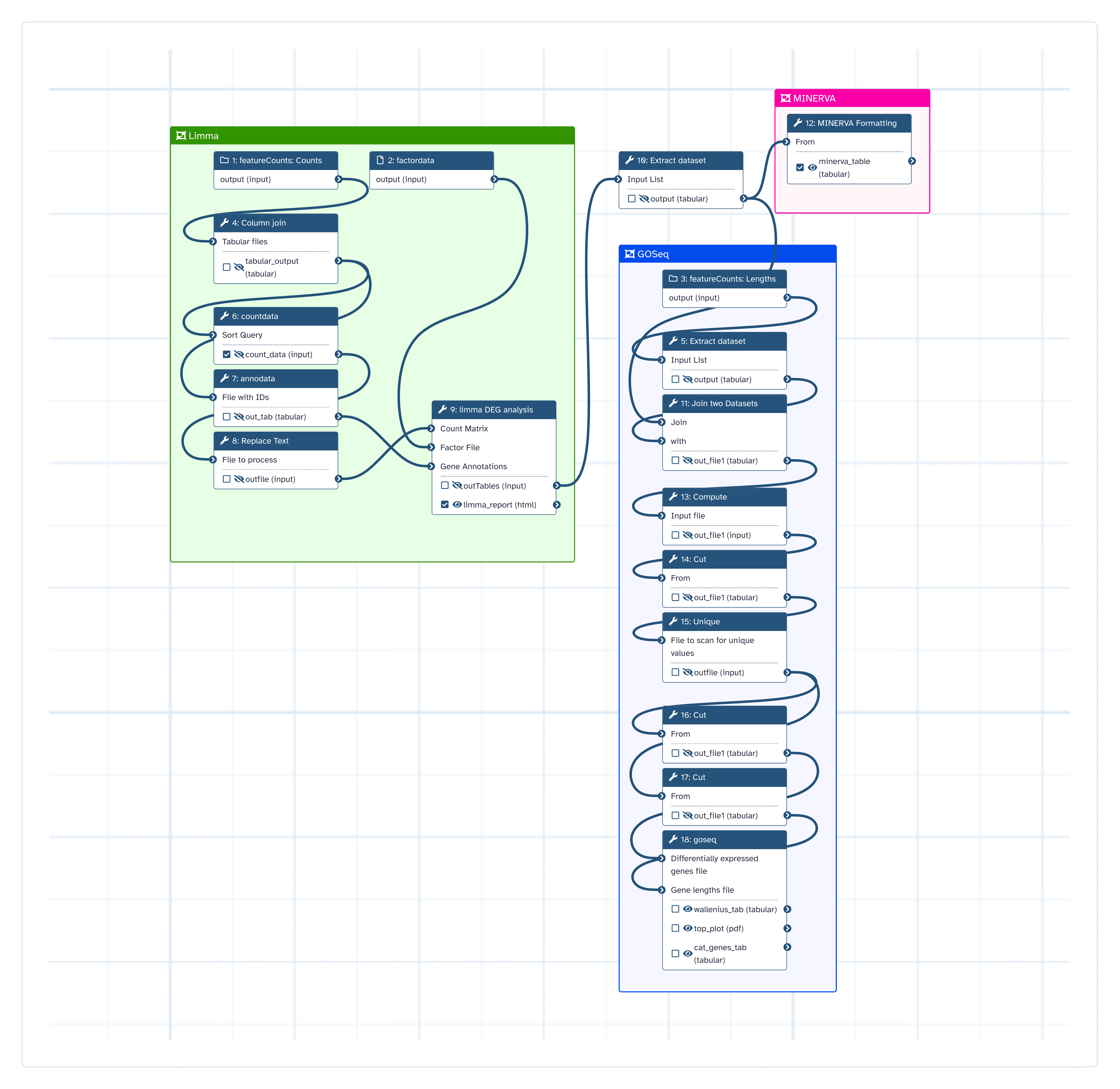
Analyse Bulk RNA-Seq data in preparation for downstream Pathways analysis with MINERVA
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| factordata | factordata | A two column factor table with (Sample Identifier, Condition) This workflow assumes a 1 factor, 2 level analysis, and was specifically designed around SARS-CoV-2 analysis with two levels, e.g. ``` SampleName Group SRR16683284 COVID SRR16683283 COVID SRR16683271 healthy SRR16683270 healthy ``` |
|
| featureCounts: Counts | featureCounts: Counts | count data collection with two column datasets (gene_id, count) |
|
| featureCounts: Lengths | featureCounts: Lengths | featureCounts Lengths collection |
|
Steps
| ID | Name | Description |
|---|---|---|
| 3 | Column join | toolshed.g2.bx.psu.edu/repos/iuc/collection_column_join/collection_column_join/0.0.3 |
| 4 | Extract dataset | __EXTRACT_DATASET__ |
| 5 | countdata | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_sort_header_tool/1.1.1 |
| 6 | annodata | toolshed.g2.bx.psu.edu/repos/iuc/annotatemyids/annotatemyids/3.16.0+galaxy1 |
| 7 | Replace Text | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_replace_in_line/1.1.2 |
| 8 | limma DEG analysis | toolshed.g2.bx.psu.edu/repos/iuc/limma_voom/limma_voom/3.50.1+galaxy0 |
| 9 | Extract dataset | __EXTRACT_DATASET__ |
| 10 | MINERVA Formatting | Cut1 |
| 11 | Join two Datasets | join1 |
| 12 | Compute | toolshed.g2.bx.psu.edu/repos/devteam/column_maker/Add_a_column1/1.4 |
| 13 | Cut | Cut1 |
| 14 | Unique | toolshed.g2.bx.psu.edu/repos/bgruening/text_processing/tp_sorted_uniq/1.1.0 |
| 15 | Cut | Cut1 |
| 16 | Cut | Cut1 |
| 17 | goseq | toolshed.g2.bx.psu.edu/repos/iuc/goseq/goseq/1.50.0+galaxy0 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| count_data | count_data | n/a |
|
| limma_report | limma_report | n/a |
|
| minerva_table | minerva_table | n/a |
|
Version History
Version 1 (earliest) Created 19th Dec 2023 at 10:10 by Helena Rasche
Initial commit
Frozen
 Version-1
Version-1b946ad9
 Creators and Submitter
Creators and SubmitterCreators
Additional credit
Clinical Bioinformatics Unit, Pathology Department, Eramus Medical Center
Submitter
Activity
Views: 4119 Downloads: 819 Runs: 0
Created: 19th Dec 2023 at 10:10
 Attributions
AttributionsNone
 Collections
Collections Run on Galaxy
Run on Galaxy https://orcid.org/0000-0001-9760-8992
https://orcid.org/0000-0001-9760-8992 Download
Download