Workflow Type: Galaxy
Checking expected species and contamination in bacterial isolate
Associated Tutorial
This workflows is part of the tutorial Checking expected species and contamination in bacterial isolate, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a Galaxy Workflow Report
Thanks to...
Workflow Author(s): Pierre, Bérénice Batut, Tristan Reynolds
Tutorial Author(s): Bérénice Batut
Tutorial Contributor(s): Clea Siguret, Tristan Reynolds, Helena Rasche, Saskia Hiltemann
Funder(s): ABRomics, The University of Melbourne, Melbourne Bioinformatics, Australian BioCommons
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| Input Paired End Reads Collection | #main/Input Paired End Reads Collection | n/a |
|
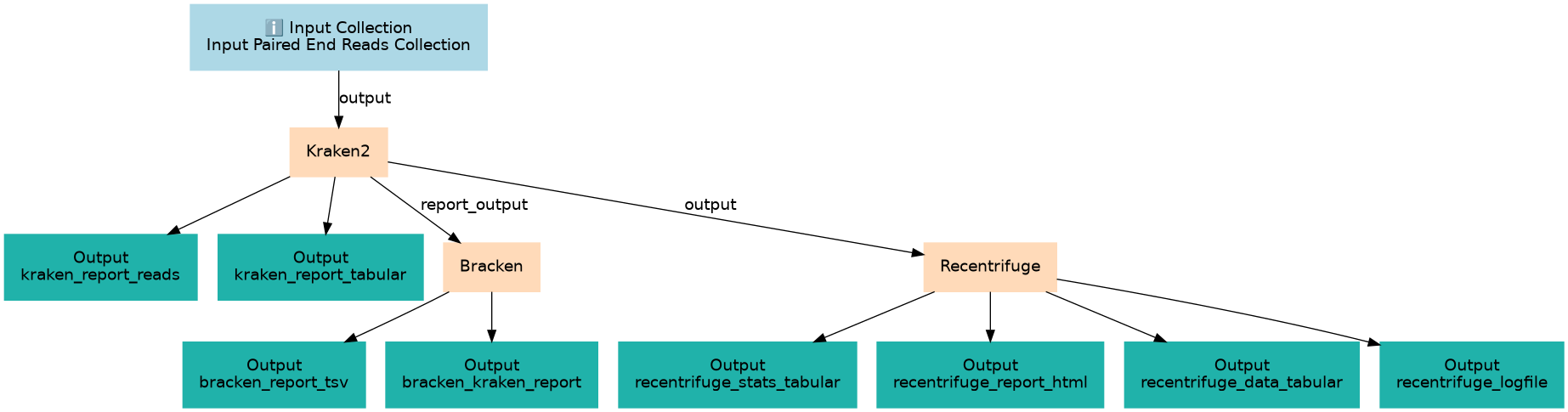
Steps
| ID | Name | Description |
|---|---|---|
| 1 | Kraken2 | toolshed.g2.bx.psu.edu/repos/iuc/kraken2/kraken2/2.1.3+galaxy2 |
| 2 | Bracken | toolshed.g2.bx.psu.edu/repos/iuc/bracken/est_abundance/3.1+galaxy0 |
| 3 | Recentrifuge | toolshed.g2.bx.psu.edu/repos/iuc/recentrifuge/recentrifuge/1.16.1+galaxy0 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| bracken_kraken_report | #main/bracken_kraken_report | n/a |
|
| bracken_report_tsv | #main/bracken_report_tsv | n/a |
|
| kraken_report_reads | #main/kraken_report_reads | n/a |
|
| kraken_report_tabular | #main/kraken_report_tabular | n/a |
|
| recentrifuge_data_tabular | #main/recentrifuge_data_tabular | n/a |
|
| recentrifuge_logfile | #main/recentrifuge_logfile | n/a |
|
| recentrifuge_report_html | #main/recentrifuge_report_html | n/a |
|
| recentrifuge_stats_tabular | #main/recentrifuge_stats_tabular | n/a |
|
Version History
 Creators and Submitter
Creators and SubmitterCreators
Not specifiedSubmitter
Discussion Channel
Tools
Activity
Views: 2591 Downloads: 460 Runs: 1
Created: 2nd Jun 2025 at 11:03
Last updated: 22nd Dec 2025 at 13:13
Annotated Properties
Scientific disciplines
Computer Science
 Attributions
AttributionsNone
 Visit source
Visit source Run on Galaxy
Run on Galaxy
 master
master 2.0
2.0