Workflow Type: Galaxy
Frozen
Frozen
Frozen
This is part of a series of workflows to annotate a genome, tagged with TSI-annotation.
These workflows are based on command-line code by Luke Silver, converted into Galaxy Australia workflows.
The workflows can be run in this order:
- Repeat masking
- RNAseq QC and read trimming
- Find transcripts
- Combine transcripts
- Extract transcripts
- Convert formats
- Fgenesh annotation
Workflow information:
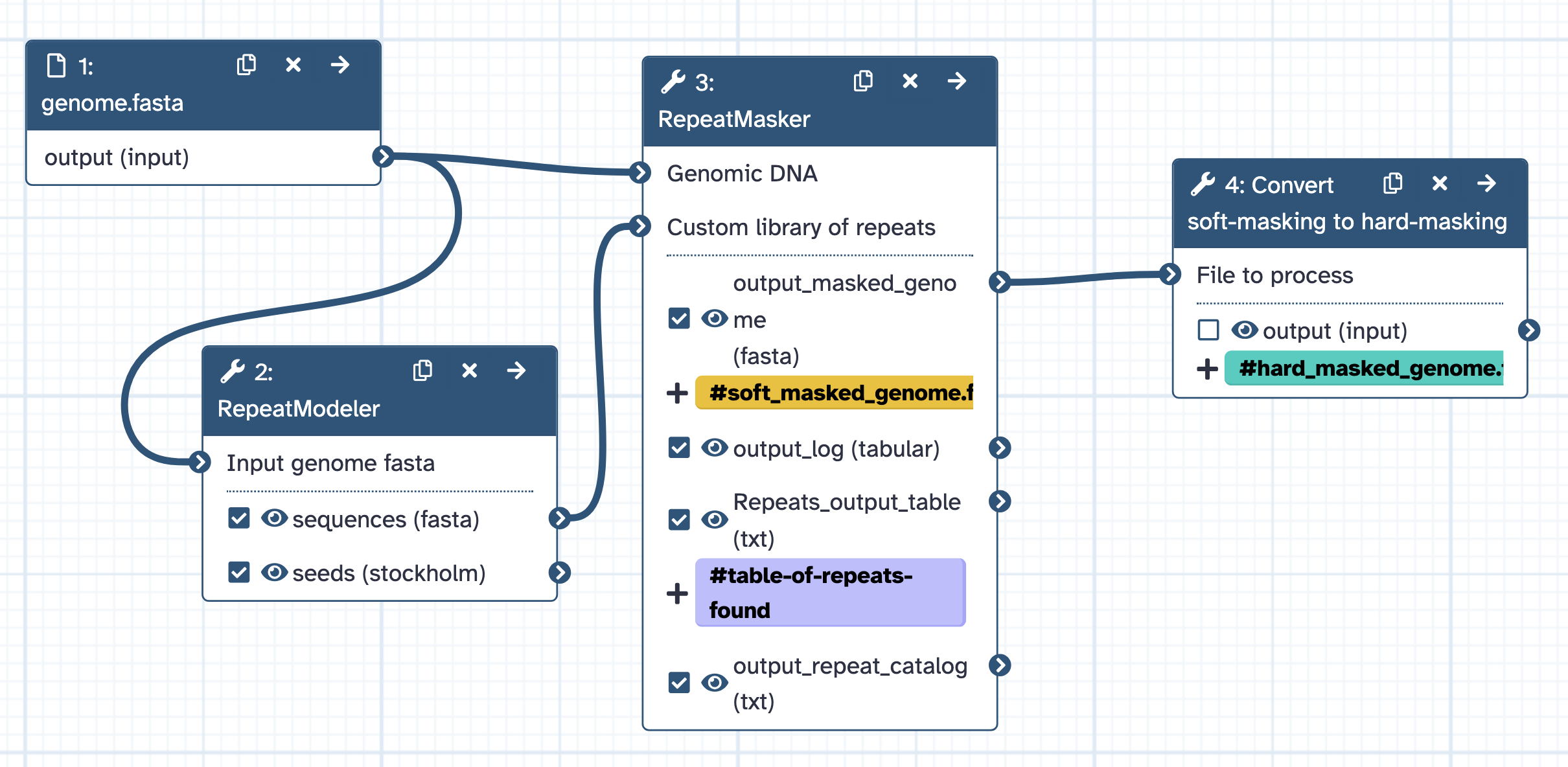
- Input = genome.fasta.
- Outputs = soft_masked_genome.fasta, hard_masked_genome.fasta, and table of repeats found.
- Runs RepeatModeler with default settings, uses the output of this (repeat library) as input into RepeatMasker.
- Runs RepeatMasker with default settings except for: Skip masking of simple tandem repeats and low complexity regions. (-nolow) : default set to yes. Perform softmasking instead of hardmasking - set to yes.
- Converts the soft-masked genome to hard-masked for for use in other tools if required.
- Workflow report displays an edited table of repeats found. Note: a known bug is that sometimes the workflow report text resets to default text. To restore, look for an earlier workflow version with correct workflow report text, and copy and paste report text into current version.
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| genome.fasta | #main/genome.fasta | n/a |
|
Steps
| ID | Name | Description |
|---|---|---|
| 1 | RepeatModeler | toolshed.g2.bx.psu.edu/repos/csbl/repeatmodeler/repeatmodeler/2.0.4+galaxy1 |
| 2 | RepeatMasker | toolshed.g2.bx.psu.edu/repos/bgruening/repeat_masker/repeatmasker_wrapper/4.1.5+galaxy0 |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| Repeats_output_table | #main/Repeats_output_table | n/a |
|
| output_log | #main/output_log | n/a |
|
| output_masked_genome | #main/output_masked_genome | n/a |
|
| output_repeat_catalog | #main/output_repeat_catalog | n/a |
|
| seeds | #main/seeds | n/a |
|
| sequences | #main/sequences | n/a |
|
Version History
Version 3 (latest) Created 21st Jun 2024 at 00:41 by Anna Syme
Added in additional step to convert soft-masking to hard-masking, for use in other tools.
Frozen
 Version-3
Version-398500e0
Version 1.1 Created 8th May 2024 at 07:24 by Anna Syme
updated to include correct workflow report text
Frozen
 Version-1.1
Version-1.188bbd9c
Version 1 (earliest) Created 8th May 2024 at 04:59 by Anna Syme
Initial commit
Frozen
 Version-1
Version-1571d961
 Creators and Submitter
Creators and SubmitterCreators
Submitter
Tools
Citation
Silver, L., & Syme, A. (2024). Repeat masking - TSI. WorkflowHub. https://doi.org/10.48546/WORKFLOWHUB.WORKFLOW.875.3
Activity
Views: 8150 Downloads: 1752 Runs: 1
Created: 8th May 2024 at 04:59
Last updated: 21st Jun 2024 at 00:45
Annotated Properties
Scientific disciplines
Biochemistry, Genetics and Molecular Biology
 Tags
Tags Attributions
AttributionsNone
 Collections
Collections Run on Galaxy
Run on Galaxy https://orcid.org/0000-0002-9906-0673
https://orcid.org/0000-0002-9906-0673