Workflow Type: Common Workflow Language
Open
Stable
Author: AMBARISH KUMAR [email protected] & [email protected]
This is a proposed standard operating procedure for genomic variant detection using GATK4.
It is hoped to be effective and useful for getting SARS-CoV-2 genome variants.
It uses Illumina RNASEQ reads and genome sequence.
Inputs
| ID | Name | Description | Type |
|---|---|---|---|
| sars_cov_2_reference_genome | n/a | n/a |
|
| rnaseq_left_reads | n/a | n/a |
|
| rnaseq_right_reads | n/a | n/a |
|
| indices_folder | n/a | n/a |
|
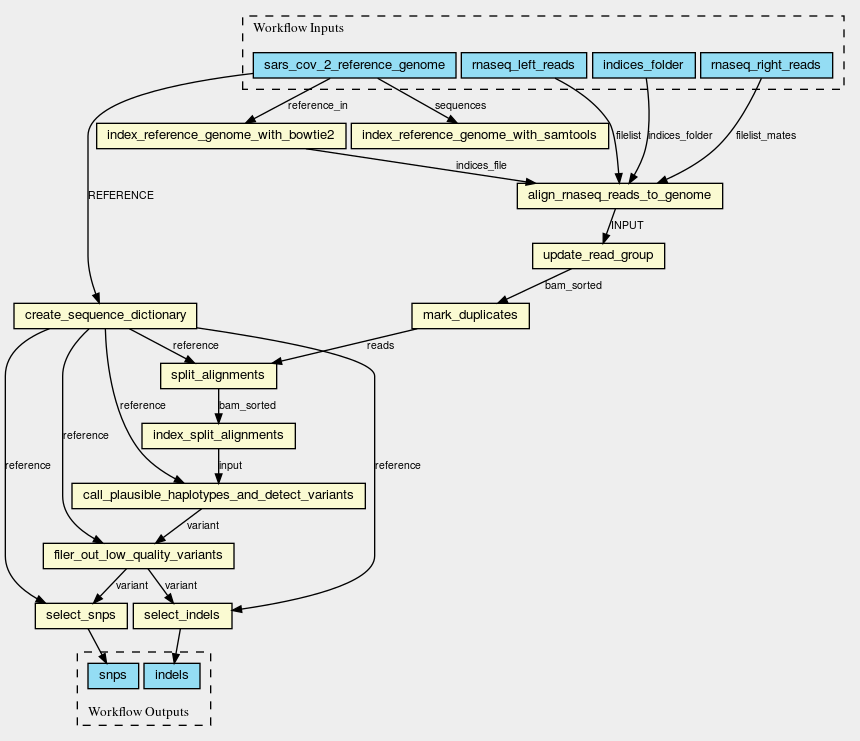
Steps
| ID | Name | Description |
|---|---|---|
| index_reference_genome_with_bowtie2 | n/a | n/a |
| align_rnaseq_reads_to_genome | n/a | n/a |
| index_reference_genome_with_samtools | n/a | n/a |
| create_sequence_dictionary | n/a | n/a |
| update_read_group | n/a | n/a |
| mark_duplicates | n/a | n/a |
| split_alignments | n/a | n/a |
| index_split_alignments | n/a | n/a |
| call_plausible_haplotypes_and_detect_variants | n/a | n/a |
| filer_out_low_quality_variants | n/a | n/a |
| select_indels | n/a | n/a |
| select_snps | n/a | n/a |
Outputs
| ID | Name | Description | Type |
|---|---|---|---|
| indels | n/a | n/a |
|
| snps | n/a | n/a |
|
Version History
Version 1 (earliest) Created 17th Jun 2020 at 07:11 by Ambarish Kumar
Added/updated 2 files
Open
 master
masterf0602cf
 Creators and Submitter
Creators and SubmitterCreator
Submitter
License
Activity
Views: 6591 Downloads: 1722
Created: 17th Jun 2020 at 07:11
Annotated Properties
Scientific disciplines
Biochemistry, Genetics and Molecular Biology
 Attributions
AttributionsNone
 View on GitHub
View on GitHub External Link
External Link